This is built with spread2() and is still experimental.
This one differs from other attempts in that it treats the advection and
dispersal as mathematical vectors that are added together.
They are "rounded" to pixel centres.
spread3(
start,
rasQuality,
rasAbundance,
advectionDir,
advectionMag,
meanDist,
dispersalKernel = "exponential",
sdDist = 1,
plot.it = 2,
minNumAgents = 50,
verbose = getOption("LandR.verbose", 0),
saveStack = NULL,
skipChecks = FALSE
)Arguments
- start
Raster indices from which to initiate dispersal
- rasQuality
A raster with habitat quality. Currently, must be scaled from 0 to 1, i.e., a probability of "settling"
- rasAbundance
A raster where each pixel represents the number of "agents" or pseudo-agents contained. This number of agents, will be spread horizontally, and distributed from each pixel that contains a non-zero non NA value.
- advectionDir
A single number or
RasterLayerin degrees from North = 0 (though it will use radians if all values areabs(advectionDir) > 2 * pi). This indicates the direction of advective forcing (i.e., wind).- advectionMag
A single number or
RasterLayerin distance units of therasQuality, e.g., meters, indicating the relative forcing that will occur. It is imposed on the total event, i.e., if themeanDistis10000, andadvectionMagis5000, then the expected distance (i.e.,63%of agents) will have settled by15000map units.- meanDist
A single number indicating the mean distance parameter in map units (not pixels), for a negative exponential distribution dispersal kernel (e.g.,
dexp). This will mean that63%of agents will have settled at thismeanDist(still experimental).- dispersalKernel
One of either
"exponential"or"weibull".- sdDist
A single number indicating the
sdparameter of a two-parameterdispersalKernel. Defaults to1, which is the same as theexponentialdistribution.- plot.it
Numeric. With increasing numbers above 0, there will be plots produced during iterations. Currently, only 0, 1, or 2+ are distinct.
- minNumAgents
Single numeric indicating the minimum number of agents to consider all dispersing finished. Default is 50.
- verbose
Numeric. With increasing numbers above 0, there will be more messages produced. Currently, only 0, 1, or 2+ are distinct.
- saveStack
If provided as a character string, it will save each iteration as part of a
rasterStackto disk upon exit.- skipChecks
Logical. If
TRUE, assertions will be skipped (faster, but could miss problems)
Value
A data.table with all information used during the spreading
Examples
## these tests are fairly heavy, so don't run during automated tests
#########################################################
# Simple case, no variation in rasQuality, numeric advectionDir and advectionMag
#########################################################
# \donttest{
library(terra)
origDTThreads <- data.table::setDTthreads(2L)
origNcpus <- options(Ncpus = 2L)
maxDim <- 10000
ras <- terra::rast(terra::ext(c(0, maxDim, 0, maxDim)), res = 100, vals = 0)
rasQuality <- terra::rast(ras)
rasQuality[] <- 1
rasAbundance <- terra::rast(rasQuality)
rasAbundance[] <- 0
# startPixel <- middlePixel(rasAbundance)
startPixel <- sample(seq(terra::ncell(rasAbundance)), 30)
rasAbundance[startPixel] <- 1000
advectionDir <- 70
advectionMag <- 4 * res(rasAbundance)[1]
meanDist <- 2600
# Test the dispersal kernel -- create a function
plotDispersalKernel <- function(out, meanAdvectionMag) {
out[, disGroup := round(distance / 100) * 100]
freqs <- out[, .N, by = "disGroup"]
freqs[, `:=`(cumSum = cumsum(N), N = N)]
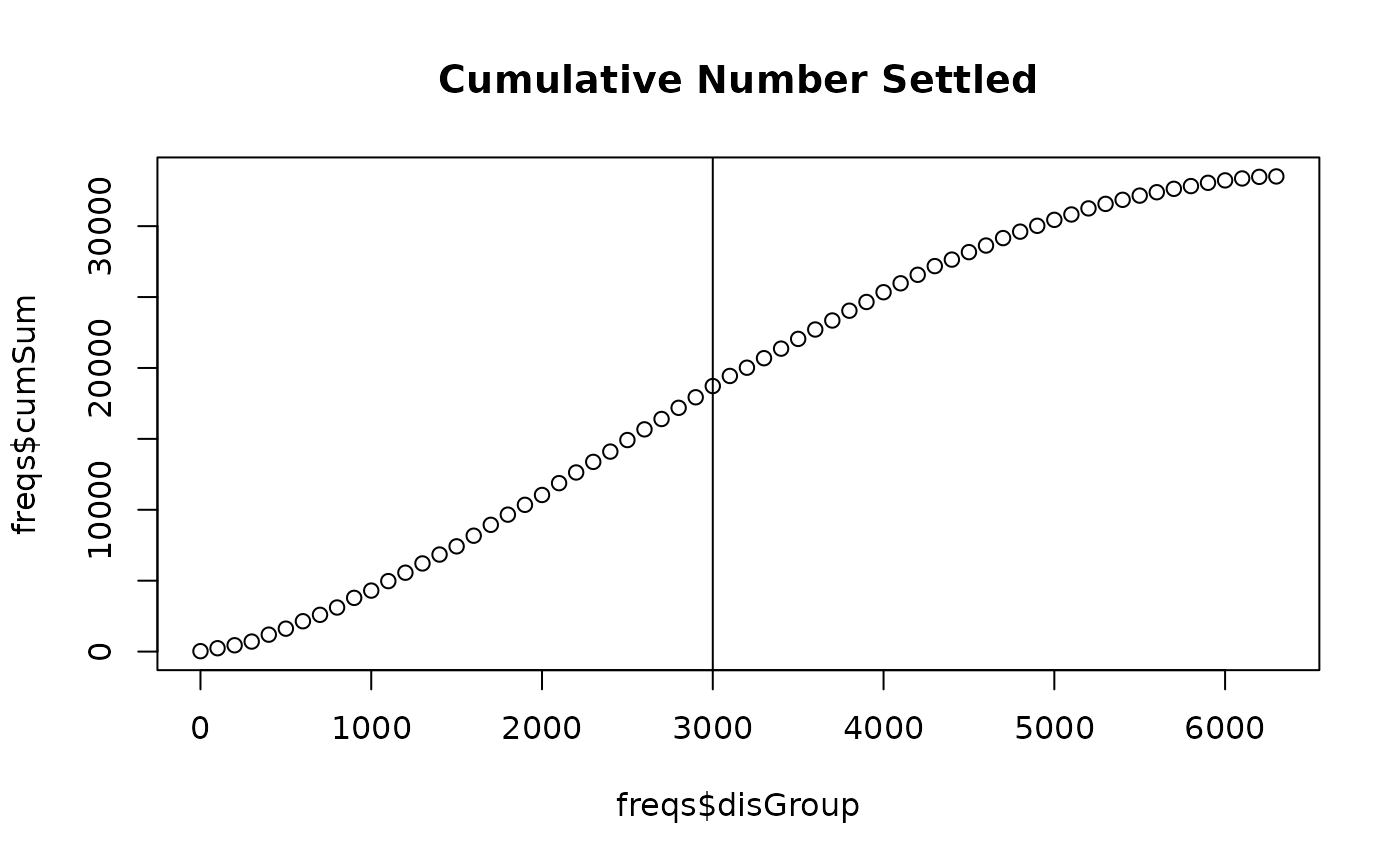
plot(freqs$disGroup, freqs$cumSum, # addTo = "CumulativeNumberSettled",
main = "Cumulative Number Settled") # can plot the distance X number
abline(v = meanAdvectionMag + meanDist)
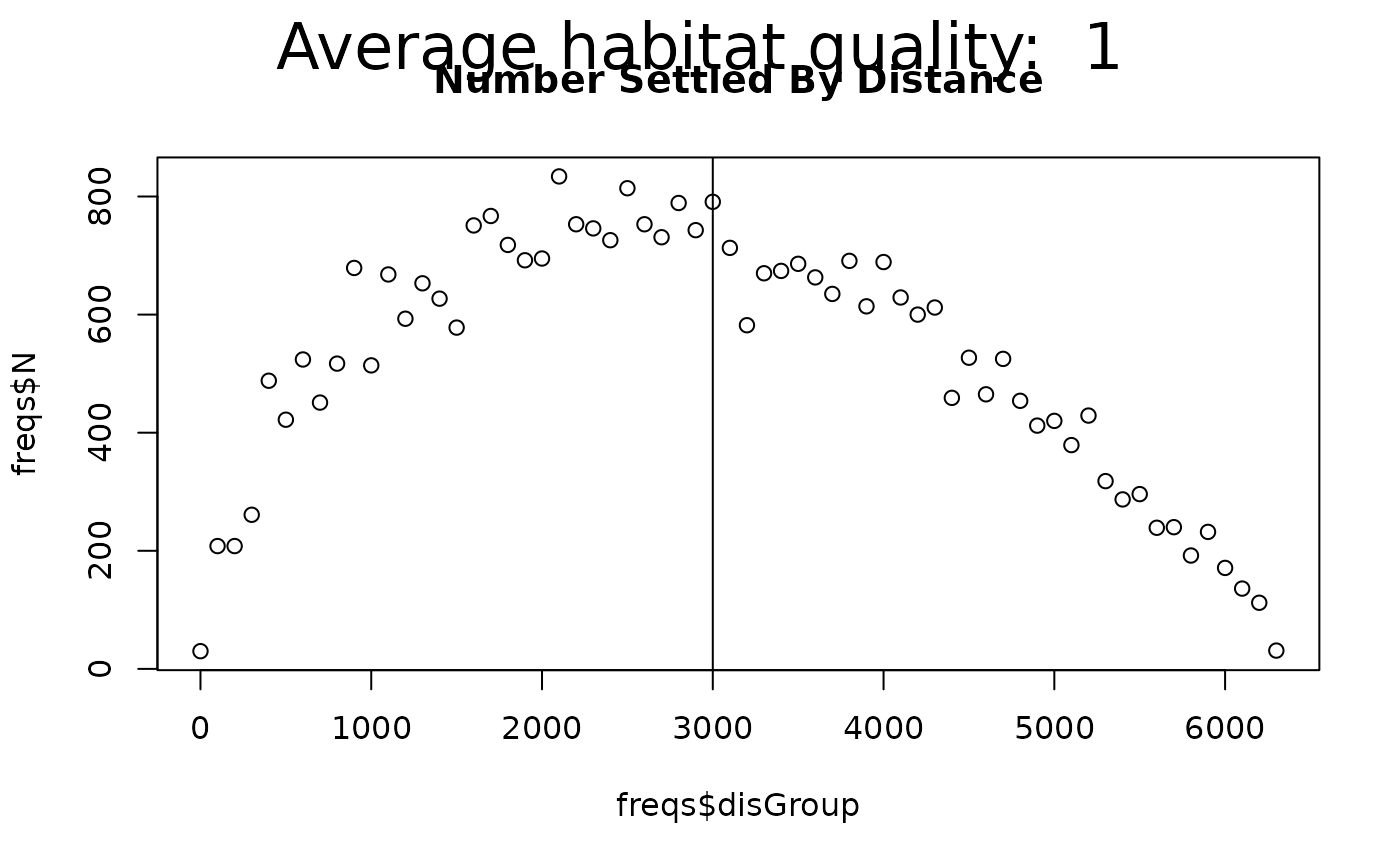
newTitle <- "Number Settled By Distance"
plot(freqs$disGroup, freqs$N, # addTo = gsub(" ", "", newTitle),
main = newTitle) # can plot the distance X number
abline(v = meanAdvectionMag + meanDist)
# should be 0.63:
freqs[disGroup == meanAdvectionMag + meanDist, cumSum] / tail(freqs, 1)[, cumSum]
mtext(side = 3, paste("Average habitat quality: ",
round(mean(rasQuality[], na.rm = TRUE), 2)),
outer = TRUE, line = -2, cex = 2)
}
out <- spread3(rasAbundance = rasAbundance,
rasQuality = rasQuality,
advectionDir = advectionDir,
advectionMag = advectionMag,
meanDist = meanDist, verbose = 2,
plot.it = interactive())
#> assuming that advectionDir is in geographic degrees(i.e., North is 0)
#> Iteration 1
#> Number still dispersing 29942.6974241127
#> Iteration 2
#> Number still dispersing 29845.4477324621
#> Iteration 3
#> Number still dispersing 29684.9978043702
#> Iteration 4
#> Number still dispersing 28993.812488292
#> Iteration 5
#> Number still dispersing 28721.1746386755
#> Iteration 6
#> Number still dispersing 28374.5983612171
#> Iteration 7
#> Number still dispersing 28013.5289421565
#> Iteration 8
#> Number still dispersing 27582.9799199363
#> Iteration 9
#> Number still dispersing 27077.3418762575
#> Iteration 10
#> Number still dispersing 25084.8618566818
#> Iteration 11
#> Number still dispersing 24119.344444795
#> Iteration 12
#> Number still dispersing 23480.4988565299
#> Iteration 13
#> Number still dispersing 22795.550955758
#> Iteration 14
#> Number still dispersing 22077.3432159583
#> Iteration 15
#> Number still dispersing 21439.5982980354
#> Iteration 16
#> Number still dispersing 20758.7939387006
#> Iteration 17
#> Number still dispersing 20069.0175900734
#> Iteration 18
#> Number still dispersing 19344.4758877966
#> Iteration 19
#> Number still dispersing 18445.5380718394
#> Iteration 20
#> Number still dispersing 17449.1183471901
#> Iteration 21
#> Number still dispersing 16598.3916918859
#> Iteration 22
#> Number still dispersing 15792.2064907562
#> Iteration 23
#> Number still dispersing 15101.8188374184
#> Iteration 24
#> Number still dispersing 14308.031009656
#> Iteration 25
#> Number still dispersing 13519.9980177877
#> Iteration 26
#> Number still dispersing 12732.4944140066
#> Iteration 27
#> Number still dispersing 11848.5250495133
#> Iteration 28
#> Number still dispersing 11168.333497928
#> Iteration 29
#> Number still dispersing 10522.2269672876
#> Iteration 30
#> Number still dispersing 9798.30085625318
#> Iteration 31
#> Number still dispersing 8821.87769414231
#> Iteration 32
#> Number still dispersing 8088.06218790952
#> Iteration 33
#> Number still dispersing 7436.56895952712
#> Iteration 34
#> Number still dispersing 6822.12286934147
#> Iteration 35
#> Number still dispersing 6252.98755623768
#> Iteration 36
#> Number still dispersing 5723.22042941179
#> Iteration 37
#> Number still dispersing 5241.41963777852
#> Iteration 38
#> Number still dispersing 4823.62593888653
#> Iteration 39
#> Number still dispersing 4402.74229009439
#> Iteration 40
#> Number still dispersing 4012.11976775426
#> Iteration 41
#> Number still dispersing 3667.66001554555
#> Iteration 42
#> Number still dispersing 3285.66271181194
#> Iteration 43
#> Number still dispersing 2966.28464160372
#> Iteration 44
#> Number still dispersing 2662.98103494232
#> Iteration 45
#> Number still dispersing 2400.59535781913
#> Iteration 46
#> Number still dispersing 2171.96351007555
#> Iteration 47
#> Number still dispersing 1931.97684779711
#> Iteration 48
#> Number still dispersing 1725.82242539007
#> Iteration 49
#> Number still dispersing 1496.94405054564
#> Iteration 50
#> Number still dispersing 1287.78294591888
#> Iteration 51
#> Number still dispersing 1142.14645025229
#> Iteration 52
#> Number still dispersing 978.596743131969
#> Iteration 53
#> Number still dispersing 858.306418743261
#> Iteration 54
#> Number still dispersing 670.567042123505
#> Iteration 55
#> Number still dispersing 600.840508414726
#> Iteration 56
#> Number still dispersing 472.722004965212
#> Iteration 57
#> Number still dispersing 422.465123908484
#> Iteration 58
#> Number still dispersing 354.608271800084
#> Iteration 59
#> Number still dispersing 265.010094370132
#> Iteration 60
#> Number still dispersing 198.784707895395
#> Iteration 61
#> Number still dispersing 175.108859796463
#> Iteration 62
#> Number still dispersing 120.988482342059
#> Iteration 63
#> Number still dispersing 32.1482342818288
plotDispersalKernel(out, advectionMag)
 # The next examples are potentially time consuming; avoid on automated testing
if (interactive()) {
#########################################################
### The case of variable quality raster
#########################################################
rasQuality <- terra::rast(system.file("extdata", "rasQuality.tif",
package = "SpaDES.tools"))
terra::crs(rasQuality) <- system.file("extdata", "targetCRS.rds", package = "SpaDES.tools") |>
readRDS() |>
slot("projargs")
mask <- rasQuality < 5
rasQuality[mask[] %in% TRUE] <- 0
# rescale so min is 0.75 and max is 1
rasQuality[] <- rasQuality[] / (reproducible::maxFn(rasQuality) * 4) + 1 / 4
rasAbundance <- terra::rast(rasQuality)
rasAbundance[] <- 0
startPixel <- sample(seq(ncell(rasAbundance)), 300)
rasAbundance[startPixel] <- 1000
advectionDir <- 75
advectionMag <- 4 * res(rasAbundance)[1]
meanDist <- 2600
out <- spread3(rasAbundance = rasAbundance,
rasQuality = rasQuality,
advectionDir = advectionDir,
advectionMag = advectionMag,
meanDist = meanDist, verbose = 2,
plot.it = interactive())
if (interactive()) {
plotDispersalKernel(out, advectionMag)
}
###############################################################################
### The case of variable quality raster, raster for advectionDir & advectionMag
###############################################################################
maxDim <- 10000
ras <- terra::rast(terra::ext(c(0, maxDim, 0, maxDim)), res = 100, vals = 0)
rasQuality <- terra::rast(ras)
rasQuality[] <- 1
rasAbundance <- terra::rast(rasQuality)
rasAbundance[] <- NA
# startPixel <- middlePixel(rasAbundance)
startPixel <- sample(seq(ncell(rasAbundance)), 25)
rasAbundance[startPixel] <- 1000
# raster for advectionDir
advectionDir <- terra::rast(system.file("extdata", "advectionDir.tif",
package = "SpaDES.tools"))
crs(advectionDir) <- crs(rasQuality)
# rescale so min is 0.75 and max is 1
advectionDir[] <- advectionDir[] / (reproducible::maxFn(advectionDir)) * 180
# raster for advectionMag
advectionMag <- terra::rast(system.file("extdata", "advectionMag.tif",
package = "SpaDES.tools"))
crs(advectionMag) <- crs(rasQuality)
# rescale so min is 0.75 and max is 1
advectionMag[] <- advectionMag[] / (reproducible::maxFn(advectionMag)) * 600
out <- spread3(rasAbundance = rasAbundance,
rasQuality = rasQuality,
advectionDir = advectionDir,
advectionMag = advectionMag,
meanDist = meanDist, verbose = 2,
plot.it = interactive())
if (interactive()) {
names(advectionDir) <- "Wind direction"
names(advectionMag) <- "Wind speed"
names(rasAbundance) <- "Initial abundances"
terra::plot(c(advectionDir, advectionMag, rasAbundance))
plotDispersalKernel(out, mean(advectionMag[]))
}
#########################################
# save iterations to a stack to make animated GIF
########################################
tmpStack <- tempfile(pattern = "stackToAnimate", fileext = ".tif")
out <- spread3(rasAbundance = rasAbundance,
rasQuality = rasQuality,
advectionDir = advectionDir,
advectionMag = advectionMag,
meanDist = 2600, verbose = 2,
plot.it = interactive(), saveStack = tmpStack)
## This animates the series of images into an animated GIF
if (require(animation, quietly = TRUE)) {
out2 <- terra::rast(tmpStack)
gifName <- file.path(tempdir(), "animation.gif")
# Only works on some systems; may need to configure
# Works on Windows without system adjustments
if (identical(.Platform$OS.type, "windows"))
saveGIF(interval = 0.1, movie.name = gifName, expr = {
for (i in seq(length(names(out2)))) terra::plot(out2[[i]])
})
}
}
# clean up
data.table::setDTthreads(origDTThreads)
options(Ncpus = origNcpus)
# }
# The next examples are potentially time consuming; avoid on automated testing
if (interactive()) {
#########################################################
### The case of variable quality raster
#########################################################
rasQuality <- terra::rast(system.file("extdata", "rasQuality.tif",
package = "SpaDES.tools"))
terra::crs(rasQuality) <- system.file("extdata", "targetCRS.rds", package = "SpaDES.tools") |>
readRDS() |>
slot("projargs")
mask <- rasQuality < 5
rasQuality[mask[] %in% TRUE] <- 0
# rescale so min is 0.75 and max is 1
rasQuality[] <- rasQuality[] / (reproducible::maxFn(rasQuality) * 4) + 1 / 4
rasAbundance <- terra::rast(rasQuality)
rasAbundance[] <- 0
startPixel <- sample(seq(ncell(rasAbundance)), 300)
rasAbundance[startPixel] <- 1000
advectionDir <- 75
advectionMag <- 4 * res(rasAbundance)[1]
meanDist <- 2600
out <- spread3(rasAbundance = rasAbundance,
rasQuality = rasQuality,
advectionDir = advectionDir,
advectionMag = advectionMag,
meanDist = meanDist, verbose = 2,
plot.it = interactive())
if (interactive()) {
plotDispersalKernel(out, advectionMag)
}
###############################################################################
### The case of variable quality raster, raster for advectionDir & advectionMag
###############################################################################
maxDim <- 10000
ras <- terra::rast(terra::ext(c(0, maxDim, 0, maxDim)), res = 100, vals = 0)
rasQuality <- terra::rast(ras)
rasQuality[] <- 1
rasAbundance <- terra::rast(rasQuality)
rasAbundance[] <- NA
# startPixel <- middlePixel(rasAbundance)
startPixel <- sample(seq(ncell(rasAbundance)), 25)
rasAbundance[startPixel] <- 1000
# raster for advectionDir
advectionDir <- terra::rast(system.file("extdata", "advectionDir.tif",
package = "SpaDES.tools"))
crs(advectionDir) <- crs(rasQuality)
# rescale so min is 0.75 and max is 1
advectionDir[] <- advectionDir[] / (reproducible::maxFn(advectionDir)) * 180
# raster for advectionMag
advectionMag <- terra::rast(system.file("extdata", "advectionMag.tif",
package = "SpaDES.tools"))
crs(advectionMag) <- crs(rasQuality)
# rescale so min is 0.75 and max is 1
advectionMag[] <- advectionMag[] / (reproducible::maxFn(advectionMag)) * 600
out <- spread3(rasAbundance = rasAbundance,
rasQuality = rasQuality,
advectionDir = advectionDir,
advectionMag = advectionMag,
meanDist = meanDist, verbose = 2,
plot.it = interactive())
if (interactive()) {
names(advectionDir) <- "Wind direction"
names(advectionMag) <- "Wind speed"
names(rasAbundance) <- "Initial abundances"
terra::plot(c(advectionDir, advectionMag, rasAbundance))
plotDispersalKernel(out, mean(advectionMag[]))
}
#########################################
# save iterations to a stack to make animated GIF
########################################
tmpStack <- tempfile(pattern = "stackToAnimate", fileext = ".tif")
out <- spread3(rasAbundance = rasAbundance,
rasQuality = rasQuality,
advectionDir = advectionDir,
advectionMag = advectionMag,
meanDist = 2600, verbose = 2,
plot.it = interactive(), saveStack = tmpStack)
## This animates the series of images into an animated GIF
if (require(animation, quietly = TRUE)) {
out2 <- terra::rast(tmpStack)
gifName <- file.path(tempdir(), "animation.gif")
# Only works on some systems; may need to configure
# Works on Windows without system adjustments
if (identical(.Platform$OS.type, "windows"))
saveGIF(interval = 0.1, movie.name = gifName, expr = {
for (i in seq(length(names(out2)))) terra::plot(out2[[i]])
})
}
}
# clean up
data.table::setDTthreads(origDTThreads)
options(Ncpus = origNcpus)
# }