Wrapper for selecting different animal movement methods.
This version uses just turn angles and step lengths to define the correlated random walk.
move(hypothesis = "crw", ...)
crw(
agent,
extent,
stepLength,
stddev,
lonlat = FALSE,
torus = FALSE,
returnMatrix = FALSE
)Arguments
- hypothesis
Character vector, length one, indicating which movement hypothesis/method to test/use. Currently defaults to 'crw' (correlated random walk) using
crw.- ...
arguments passed to the function in
hypothesis- agent
A
SpatVectorpoints geometry or aSpatialPoints*(deprecated) object. If is has attributes, e.g.,SpatialPointsDataFrame, 2 of the columns must bex1andy1, indicating the previous location. If it does not have these columns as attributes,x1andy1will be assigned randomly.- extent
An optional
Extentobject that will be used fortorus.- stepLength
Numeric vector of length 1 or number of agents describing step length.
- stddev
Numeric vector of length 1 or number of agents describing standard deviation of wrapped normal turn angles.
- lonlat
Logical. If
TRUE, coordinates should be in degrees. IfFALSEcoordinates represent planar ('Euclidean') space (e.g. units of meters)- torus
Logical. Should the movement be wrapped to the opposite side of the map, as determined by the
extentargument. DefaultFALSE.- returnMatrix
If
TRUEthen the return object will be amatrix. This will be MUCH faster than retaining thesporSpatVectorclass, and thus will be much more effective for iterativecrwcalls
Value
A SpatVector points object with updated spatial position defined
by a single occurrence of step length(s) and turn angle(s).
Details
This simple version of a correlated random walk is largely the version that was presented in Turchin 1998, but it was also used with bias modifications in McIntire, Schultz, Crone 2007.
References
Turchin, P. 1998. Quantitative analysis of movement: measuring and modeling population redistribution in animals and plants. Sinauer Associates, Sunderland, MA.
McIntire, E. J. B., C. B. Schultz, and E. E. Crone. 2007. Designing a network for butterfly habitat restoration: where individuals, populations and landscapes interact. Journal of Applied Ecology 44:725-736.
See also
Examples
origDTThreads <- data.table::setDTthreads(2L)
origNcpus <- options(Ncpus = 2L)
# using just matrix
N <- 10
xrange <- yrange <- c(-50, 50)
starts <- cbind(x = stats::runif(N, xrange[1], xrange[2]),
y = stats::runif(N, yrange[1], yrange[2]))
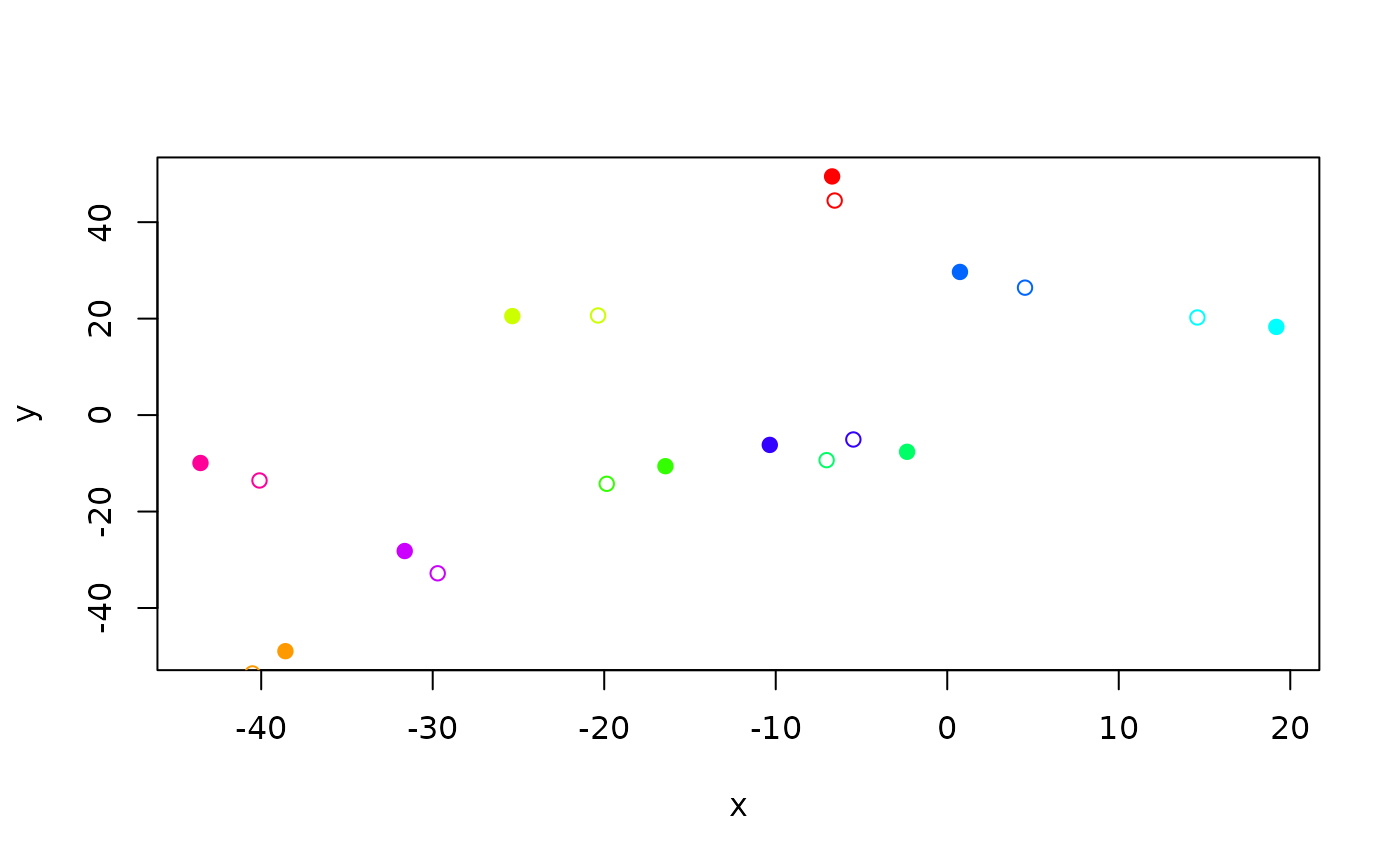
moved <- crw(starts, stepLength = 5, stddev = 10)
plot(starts, col = rainbow(10), pch = 19)
points(moved, col = rainbow(10))
 # as SpatVector
agent <- terra::vect(starts)
moved <- crw(agent, stepLength = 5, stddev = 10)
movedAgain <- crw(moved, stepLength = 5, stddev = 10)
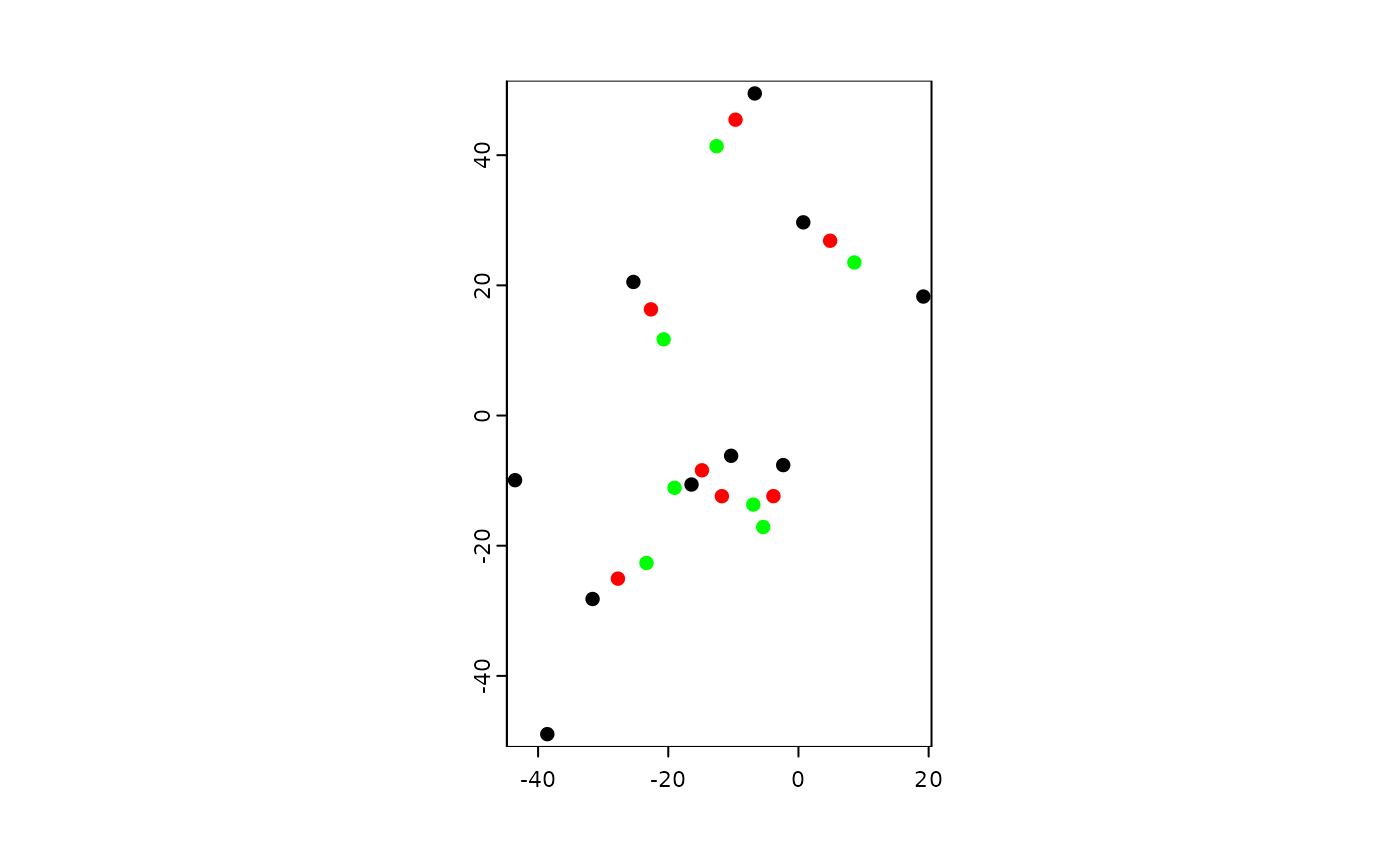
terra::plot(agent)
terra::plot(moved, add = TRUE, col = "red")
terra::plot(movedAgain, add = TRUE, col = "green")
# as SpatVector
agent <- terra::vect(starts)
moved <- crw(agent, stepLength = 5, stddev = 10)
movedAgain <- crw(moved, stepLength = 5, stddev = 10)
terra::plot(agent)
terra::plot(moved, add = TRUE, col = "red")
terra::plot(movedAgain, add = TRUE, col = "green")
 # 1000x faster!! -- returnMatrix = TRUE
agentOrig <- agent
reps <- 1e2
system.time({
for (i in 1:reps) agent <- crw(agent, stepLength = 5, stddev = 10, returnMatrix = TRUE)
})
#> user system elapsed
#> 0.009 0.000 0.008
agent <- agentOrig
system.time({
for (i in 1:reps) agent <- crw(agent, stepLength = 5, stddev = 10)
})
#> user system elapsed
#> 0.849 0.000 0.849
# as sp
if (requireNamespace("sp")) {
agent <- sp::SpatialPoints(starts)
spdf <- crw(agent, stepLength = 5, stddev = 10)
spdfNew <- crw(spdf, stepLength = 5, stddev = 10)
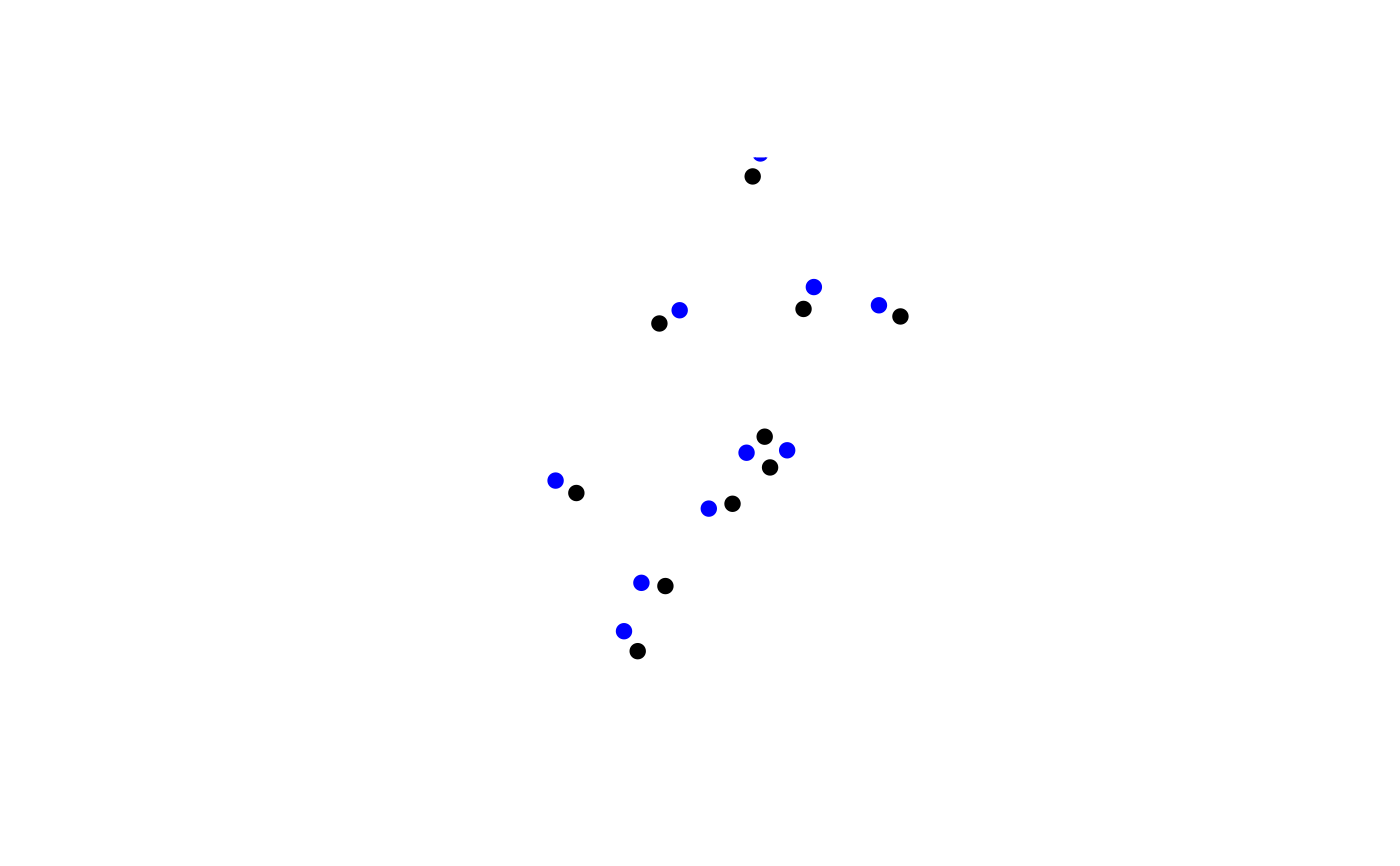
terra::plot(spdf, pch = 19)
terra::points(spdfNew, col = "blue", pch = 19)
}
#> agent does not have columns named x1 and y1, which represent the 'previous' locations. Assigning random values to those columns.
#> agent does not have columns named x1 and y1, which represent the 'previous' locations. Assigning random values to those columns.
# 1000x faster!! -- returnMatrix = TRUE
agentOrig <- agent
reps <- 1e2
system.time({
for (i in 1:reps) agent <- crw(agent, stepLength = 5, stddev = 10, returnMatrix = TRUE)
})
#> user system elapsed
#> 0.009 0.000 0.008
agent <- agentOrig
system.time({
for (i in 1:reps) agent <- crw(agent, stepLength = 5, stddev = 10)
})
#> user system elapsed
#> 0.849 0.000 0.849
# as sp
if (requireNamespace("sp")) {
agent <- sp::SpatialPoints(starts)
spdf <- crw(agent, stepLength = 5, stddev = 10)
spdfNew <- crw(spdf, stepLength = 5, stddev = 10)
terra::plot(spdf, pch = 19)
terra::points(spdfNew, col = "blue", pch = 19)
}
#> agent does not have columns named x1 and y1, which represent the 'previous' locations. Assigning random values to those columns.
#> agent does not have columns named x1 and y1, which represent the 'previous' locations. Assigning random values to those columns.
 # clean up
data.table::setDTthreads(origDTThreads)
options(Ncpus = origNcpus)
# clean up
data.table::setDTthreads(origDTThreads)
options(Ncpus = origNcpus)